Monday, June 1, 2020
Submissions Due June 8
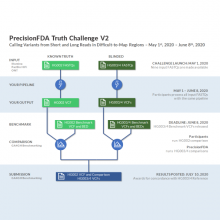
The last day to submit to precisionFDA’s Truth Challenge V2: Calling Variants from Short and Long Reads in Difficult-to-Map Regions is June 8.
Join this challenge to assess variant calling pipeline performance on a common frame of reference consisting of difficult-to-map regions, segmental duplications, and the Major Histocompatibility Complex (MHC). Ilumina, PacBio HiFi, and Oxford Nanopore sequencing datasets will be made available for this challenge.
In the context of whole human genome sequencing, software pipelines typically rely on mapping sequencing reads or assemblies to a reference genome and the subsequent identification of variants. One way of assessing the performance of such pipelines is by using well-characterized reference datasets such as Genome in a Bottle’s 7 human genome benchmarks. Two of these benchmarks were used in the first precisionFDA Truth Challenge, which assessed variant calling accuracy in “easier-to-map” regions of the genome. The Genome in a Bottle Consortium led by the National Institute of Standards and Technology (NIST) has developed expanded benchmarks for HG003 and HG004, the parents of an Ashkenazi trio. Before their release, precisionFDA and NIST are running a new truth challenge focused on variant calling in more challenging genomic regions.
For more information, visit the challenge site here. Participants are required to register for a precisionFDA user account.


