June Data Sharing and Reuse Seminar
Friday, June 12, 2026

Kevin J. Lane, Ph.D., M.A. will present "CAFE: Data-sharing and Resources to Accelerate Health and Extreme Weather Research" and Kyle P. Messier, Ph.D. will present "Geospatial Tools for Large Scale Environmental and Human Health Analyses" from 12:00 p.m.–1:00 p.m. EDT.
About the Seminars
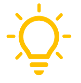
The CAFE supports the health and extreme weather (HEW) research community of practice by facilitating data sharing, standardization, and reuse through centralized infrastructure and tools. A primary accomplishment is the expansion of the CAFE Dataverse repository, now hosting over 1,200 datasets including the successful migration of 300+ SEDAC datasets along with the creation of new research group sub-collections and enhanced tracking of data use and impact. The DM team has also developed reproducible workflows and open-source tools (e.g., GitHub tutorials for ERA5 data aggregation) to enable standardized data processing across global contexts. In parallel, CAFE has advanced harmonization efforts through metadata standardization, data ontologies, and community-informed data needs assessments, exemplified by a recently published survey of HEW data practices. Ongoing initiatives focus on expanding dataset ingestion and integrating.
Large scale geospatial data such as satellite imagery, land cover, meteorology, and geospatial models are incredibly useful and powerful for understanding the impacts of the environment on human health. However, the data are notoriously difficult to use. The Spatiotemporal Exposure and Toxicology group (SET) group, at the National Institute of Environmental Health Sciences, has developed open-source tools to aide in this process and lower the burden of expertise on geospatial computations needed to conduct environmental health analyses. In this talk, I will discuss the development and applications of the open-source R-packages for integrating environmental data into health analyses.
About the Speakers
Kevin J. Lane, PhD, MA, is an Associate Professor of Environmental Health at the Boston University School of Public Health. His research focuses on the health impacts of air pollution, urbanization, and extreme weather across local, national, and global contexts. Leveraging expertise in geospatial science, big data, and exposure assessment, he develops innovative methods that integrate GIS, remote sensing, and time-activity data to advance environmental epidemiology. Dr. Lane is Director of the Environmental Health doctoral program and Associate Director of the BUSPH Center for Health Data Science, and he contributes to multiple national research initiatives to enhance data-sharing, including the NIEHS-funded CAFE program and the NIA Gateway Exposome Coordinating Center. His work spans diverse topics such as aviation-related air pollution, urban heat, and climate-health interactions, with applications to outcomes including cardiovascular disease, birth outcomes, and mortality.
The research of Kyle P. Messier, Ph.D. is fundamentally based in spatial and spatiotemporal statistical methods and applications. The common theme is spatial statistics methods and applications in population-level exposure and risk assessment. Messier is a Stadtman Tenure-Track Investigator at the National Institute of Environmental Health Sciences (NIEHS) in the Division of Translational Toxicology (DTT). He leads the Spatiotemporal Exposures and Toxicology group {S.E.T.} within the Predictive Toxicology Branch. He also holds a joint appointment with the National Institute of Minority Health and Health Disparities (NIMHD) in Bethesda, Maryland and a secondary appointment in the NIEHS Biostatistics and Computational Biology Branch. Messier received a B.S. in Environmental Studies from the University of North Carolina at Asheville and a M.S. and Ph.D. from the University of North Carolina at Chapel Hill.
About the Seminar Series
The seminar is open to the public and registration is required each month. Individuals who need interpreting services and/or other reasonable accommodations to participate in this event should contact Allison Hurst at 301-670-4990. Requests should be made at least five days in advance of the event.
The National Institutes of Health (NIH) Office of Data Science Strategy hosts this seminar series to highlight examples of data sharing and reuse on the second Friday of each month at noon ET. The monthly series highlights researchers who have taken existing data and found clever ways to reuse the data or generate new findings. A different NIH institute or center will also share its data science activities each month.